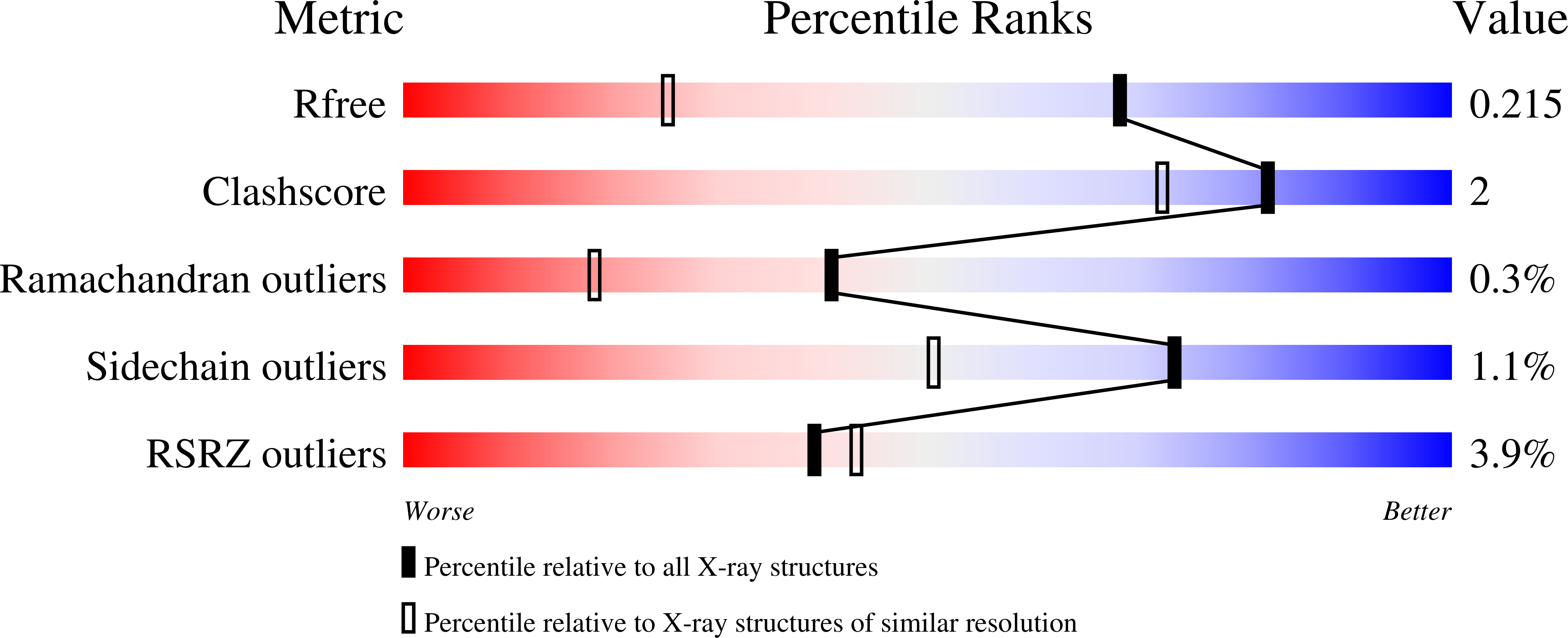
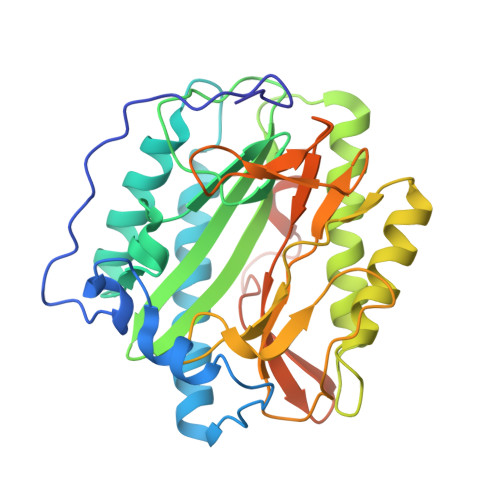
Structural analysis of bengamide derivatives as inhibitors of methionine aminopeptidases.
Xu, W., Lu, J.P., Ye, Q.Z.(2012) J Med Chem 55: 8021-8027
- PubMed: 22913487
- DOI: https://doi.org/10.1021/jm3008695
- Primary Citation of Related Structures:
4FLI, 4FLJ, 4FLK, 4FLL - PubMed Abstract:
Natural-product-derived bengamides possess potent antiproliferative activity and target human methionine aminopeptidases (MetAPs) for their cellular effects. Several derivatives were designed, synthesized, and evaluated as MetAP inhibitors. Here, we present four new X-ray structures of human MetAP1 in complex with the inhibitors. Together with the previous structures of bengamide derivatives with human MetAP2 and tubercular MtMetAP1c, analysis of the interactions of these inhibitors at the active site provides structural basis for further modification of these bengamide inhibitors for improved potency and selectivity as anticancer and antibacterial therapeutics.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, Indiana University School of Medicine, Indianapolis, Indiana 46202, USA.